Make your models shine!
References
- Jinkō - The Modeling and Trial Simulation Platform: https://www.jinko.ai
- API & Platform Documentation - Key features, REST API and scientific documentation: https://doc.jinko.ai/api/#/
- Github: API Helpers - Python helpers function to ease use of Jinko's API: https://github.com/novainsilico/jinko-api-helpers-python
- Github: Cookbooks - Practical examples and tutorials for using the Jinko API: https://github.com/novainsilico/jinko-api-cookbook
Workshop Series: Empowering Modelers with Jinkō
Program structure
- Each workshop will last 2 hours, consisting of 30 to 45 minutes of instruction and demo and the remainder as hands-on practice.
- Workshops can be taken in any order (beware though of non mandatory prerequisites) allowing flexibility for participants to tailor their learning path.
- Materials provided: Workshop-specific assets, datasets, and access to a demo Jinkō environment with a Jinkō account.
This structure provides a comprehensive training program, addressing both technical and collaborative aspects of Jinkō’s platform for modelers.
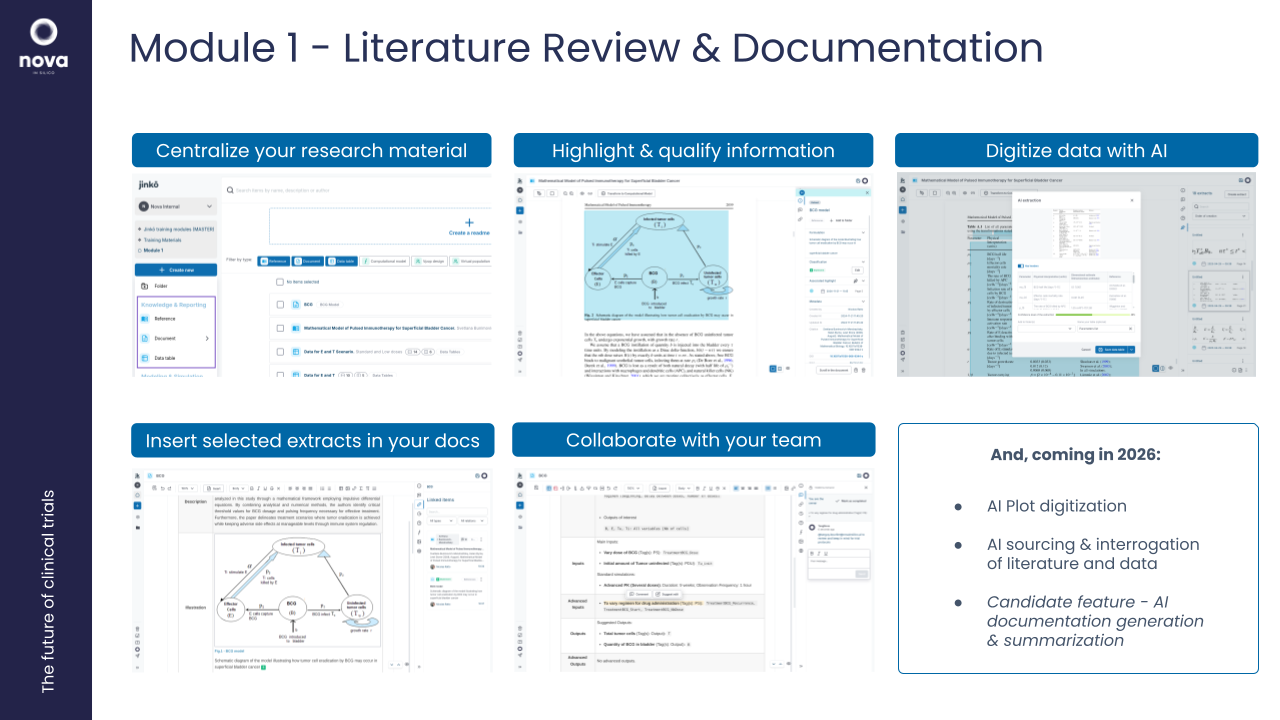
1. Workshop: Literature Review & AI-Assisted Knowledge Extraction in Jinkō
Objective: Help participants efficiently extract, organize, and utilize knowledge from the literature using Jinkō’s AI tools.
Overview:
- Introduction to the Jinkō knowledge system for capturing scientific data.
- Using the AI tools for extraction
- Best practices for systematic and transparent knowledge and data extraction with traceability.
Key Topics:
- Managing and organizing references.
- Using AI to extract equation and data
- Highlight key information and insert into the knowledge repository.
- Creating a collaborative knowledge database for model building.
Interactive Component:
- Participants will perform a mock literature extraction using a predefined set of publications relevant to their projects.

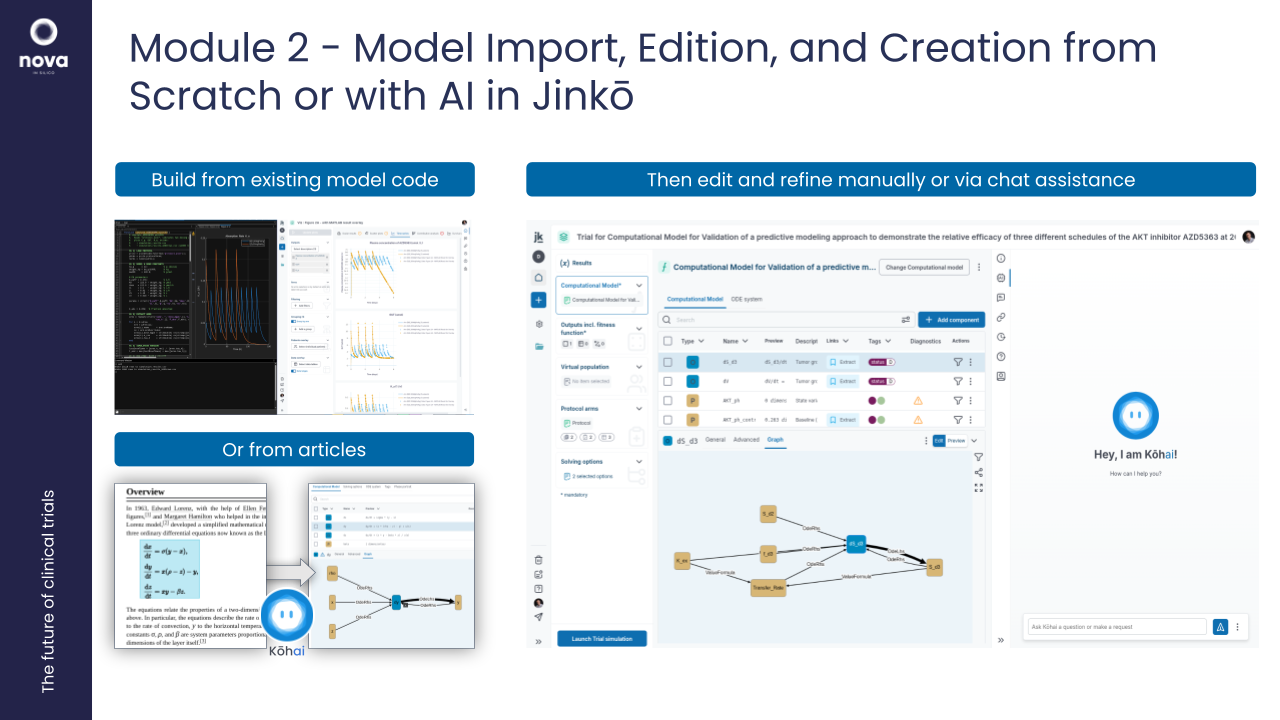
2. Workshop: Model Import, Edition, and Creation from Scratch or with AI in Jinkō
Objective: Equip participants with the skills to upload, edit, and create models in Jinkō, using its intuitive model-building tools.
Overview:
- Uploading models (e.g., SBML or Simbiology format) into Jinkō.
- Editing and documenting existing models or building new ones from scratch.
- Visualizing and managing model components through dynamic representations.
- Creating a model from paper using AI
Key Topics:
- Autocompletion of parameters, annotations, and unit checks.
- Step-by-step guide on using tags for faster model use downstream.
Interactive Component:
- Hands-on session where participants import and edit a basic pharmacokinetics (PK) model.

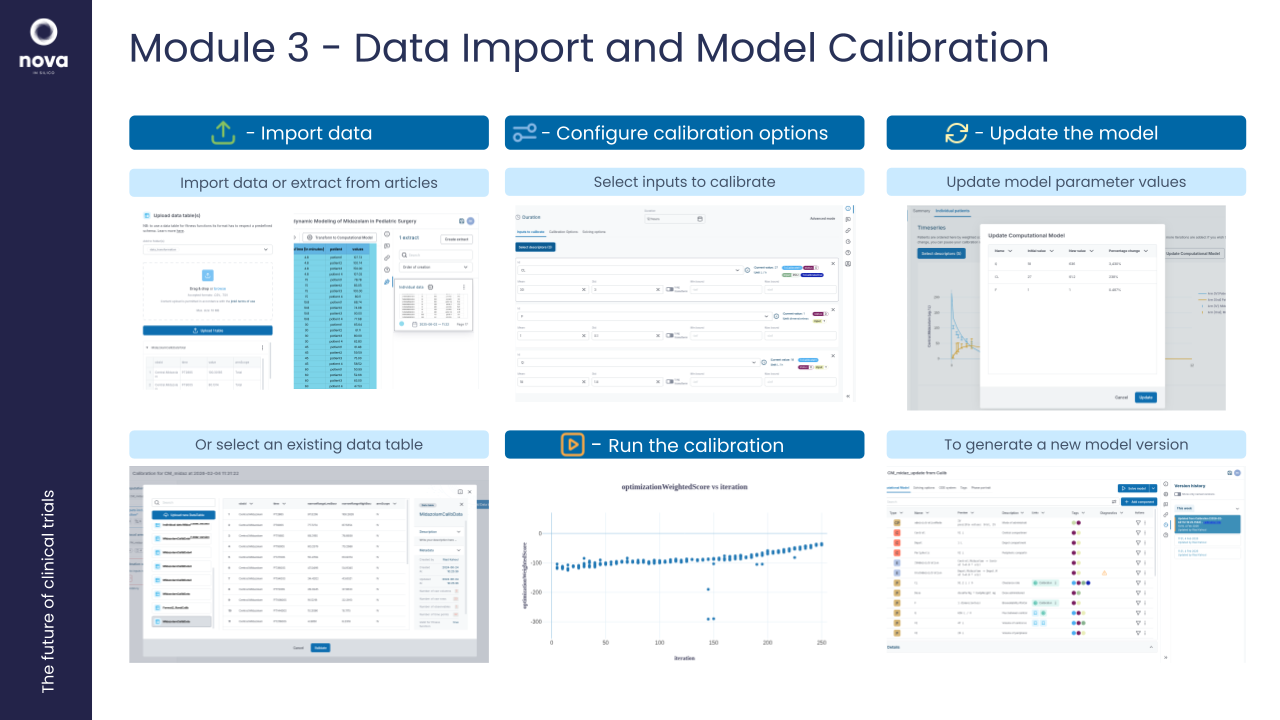
3. Workshop: Data Import & Model Calibration in Jinkō
Objective: Teach participants how to bring external data into Jinkō, and calibrate models to ensure they accurately reflect real-world biology and clinical outcomes.
Overview:
- Importing structured (clinical trial or lab) data
- Formalized and unstructured data (literature, qualitative insights) into scorings
- Parametrizing models with genetic optimization algorithms (CMAES) and fitting virtual populations.
Key Topics:
- Best practices for data handling and model calibration.
Interactive Component:
- Participants will import a dataset, calibrate a simple model, simulate and sub-sample a virtual population.
Recommended prerequisite: Workshop 1 and 2
4. Workshop: Running Clinical Trials in Jinkō
Objective: Introduce participants to the process of setting up, simulating, and analyzing virtual clinical trials using Jinkō.
Overview:
- Introduction to trial simulations: setting up protocol arms, adjusting doses, and adding variability via a virtual population.
- Running scalable simulations and tracking progress in real-time.
- Population calibration via sub-sampling of completed trial results
Key Topics:
- Trial design and optimization.
- Result visualization: Kaplan-Meier survival curves, hazard rates, and endpoint analysis.
- Creating and fine-tuning virtual populations for clinical trial simulations.
Interactive Component:
- Simulation of a simplified phase III trial and comparison of trial arms.
Recommended prerequisite: Workshop 2

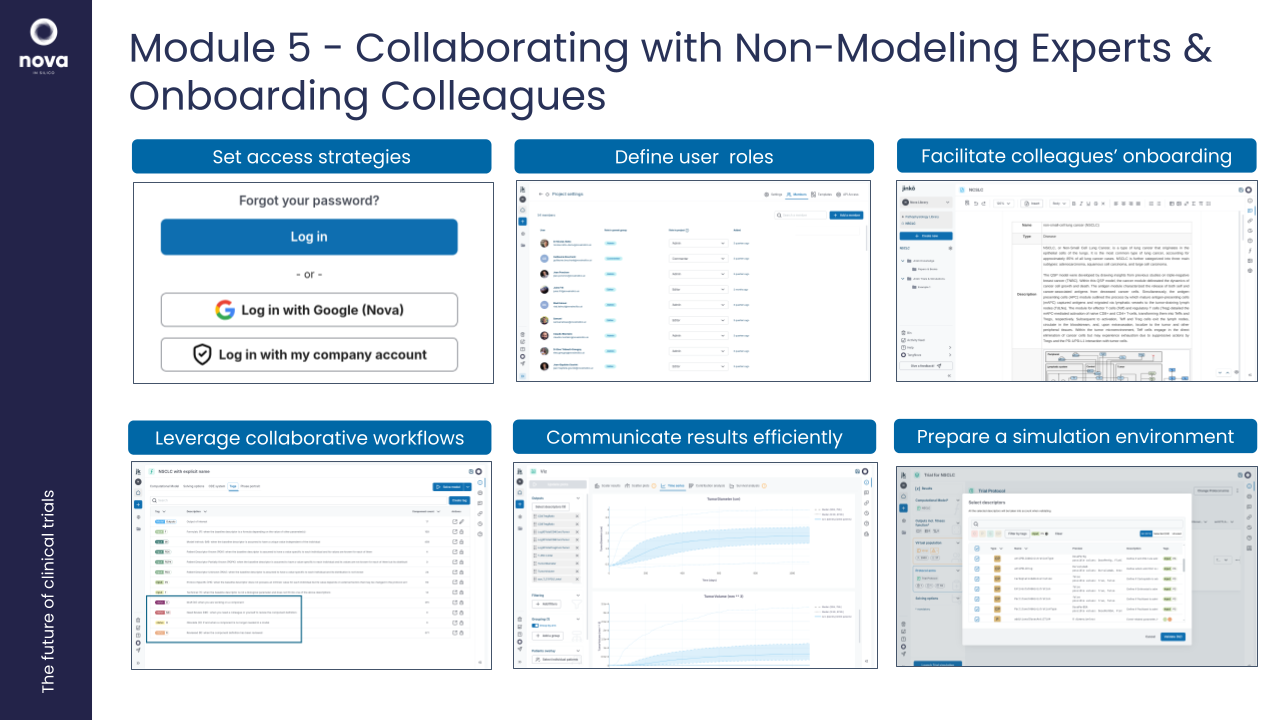
5. Workshop: Collaborating with Non-Modeling Experts & Onboarding Colleagues
Objective: Facilitate interdisciplinary collaboration and streamline the onboarding process for teams not expert in modeling.
Overview:
- How to present and document models for broad accessibility and safety.
- Collaborative features in Jinkō: shared model annotations, comments, and audits.
- Communicating results effectively to non-modeling stakeholders.
- Prepare a simulation & analysis environment for non-modeling stakeholders.
Key Topics:
- Presenting complex models for internal and external team presentations.
- Collaborative workflows in Jinkō for interdisciplinary teams.
Interactive Component:
- Participants will create a shared project, assign roles, and collaboratively edit, run and analyze a trial.
Recommended prerequisite: Workshop 4

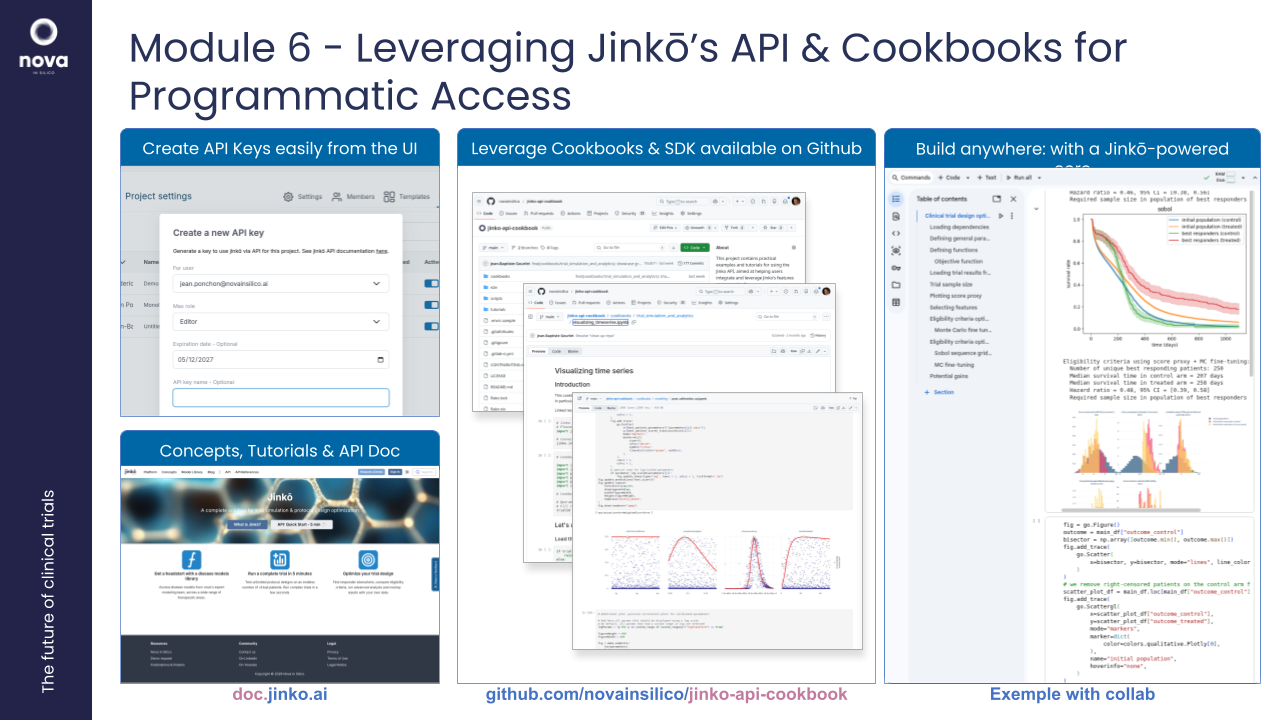
6. Workshop: Leveraging Jinkō’s API & Cookbooks for Programmatic Access
Objective: Enable participants to use the Jinkō API for more flexible, automated workflows, integrating models with external tools.
Overview:
- Overview of Jinkō’s API and Jinko SDK for Python
- Introduction to the open source cookbooks for practical examples of API use.
Key Topics:
- Using the API to programmatically manage trial resources (disease model, population, protocols, ..), launch simulations and analyze results.
- Integrating Jinkō with external environments like machine learning pipelines or visualization tools.
Interactive Component:
- Set up a Jupyter-based workbench and an API key to interact with a project on Jinko
- Learn by doing: create a population and a trial, launch a distributed simulation, analyze and visualize results.
- Play with practical examples of real pipelines & integration.
Recommended prerequisite: Workshop 4
